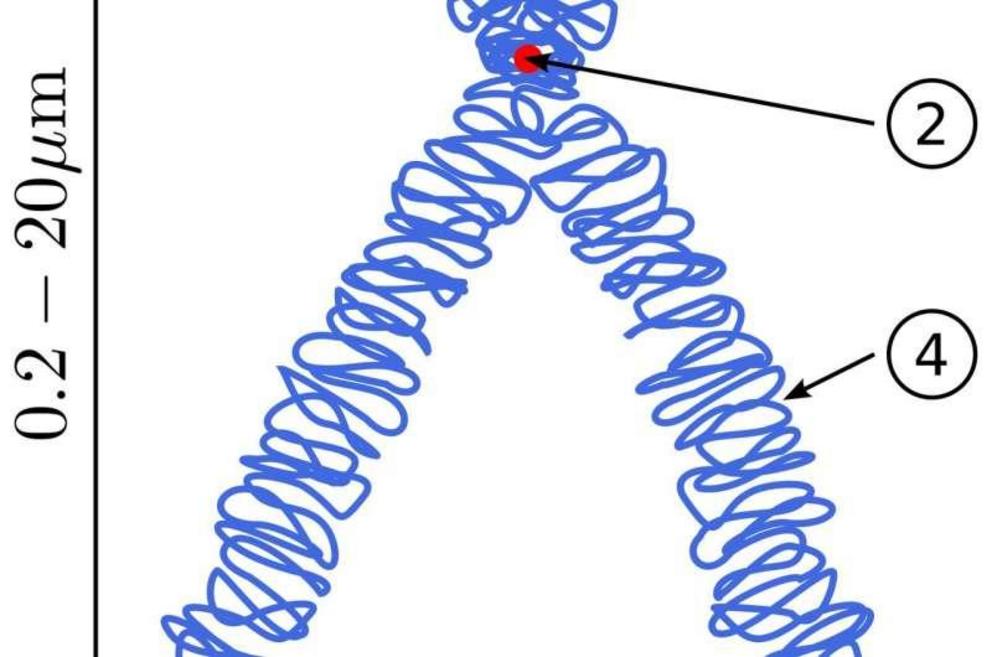
3-D creature construction: building chromosomes from the ground up
The Genome in a Box project is the brainchild of researchers Anthony Birnie and Cees Dekker from the Dept. of Bionanoscience at the Delft University of Technology. Their stated goal is to assemble a functioning chromosome from the bottom-up, beginning with the naked DNA. In theory, the raw sequence could be printed in pieces using DNA synthesis machines and then stitched together into one long string with the correct code of the desired chromosome. That would be nearly impossible in practice, at least with our existing technology. There would be no way to keep the fragile strings sorted so that they could be properly joined and folded without error.
Instead, what the researchers propose to do is to take a deproteinated genome-sized DNA string isolated from live cells, and then carefully add back appropriate DNA organizing elements one by one to bind and compact the sequence into a proper chromosome within some suitably advanced microfluidics apparatus. These DNA organizers include various histone proteins that wind up individual ~200 base pair segments of DNA to form a condensed beads-on-a-string configuration. These nucleosomes are then bridged together and extruded into higher-order loops by SMC (structural maintenance of chromosomes) motor proteins. Typical SMCs, like cohesin and condensin complexes, can rapidly reel in DNA at a rate of 2000 bps/sec, but can only pull with a stall force of about a piconewton.
These loops are then further handcuffed together to form large, topologically associating domains (TADs), which, at least for mammals, are generated on a scale of about 880 kb. Epigenetic markers are also added to define transcriptionally active (euchromatin) or repressed (heterochromatin) domains, which are, in turn, ultimately organized into distinct chromosome territories within the nucleus. Other proteins, including polymerases, track along the chain, introducing positive supercoiling and torsional stress ahead of them, and negative supercoils in their wake, while topoisomerases help decatenate interwoven chains to further control topology.
In their recent article for ACS Nano, the author's style their forward-thinking genome-in-a-box plans after the so-called particle-in-a-box theoretical construct so valued in physics. They further propose that overarching physical principles, including phase separation and ideas from polymer physics, can help guide the experiment. Their hope is that these ground-floor in vitro studies will give insight into the true nature of chromosomes that the currently used top-down, fluorescence-based imaging and immunoprecipitation methods simply can not give. But will all this actually work? In naturally occurring chromosomes, for instance, the above processes do not always follow each other in a lock-step sequence. Epigenetic adjustments and histone wrapping could be happening in one spot, while loop extrusion and supercoiling happening in another. In fact, all the above could be occurring in one form or another simultaneously.
While the goal of recreating a full eukaryotic, or even bacterially derived chromosome is a noble effort, perhaps scienctists could begin with something a little more tractable. The mitochondrial DNA (mtDNA) is organized into a nucleoid of only about 16,600 bps. By contrast, even our smallest chromosome, chromosome 21 (not 22 or Y) is several orders of magnitude larger, at 48 million bps. In fact, efforts to recreate artificial nucleoids in vitro have already had some initial success using just a few organizing proteins. For example, a minimal mitochondrial replisome was used to establish that Twinkle is the helicase used at the mtDNA replication fork. The definitive mtDNA polymerase (POLγ) cannot use double-stranded DNA template for DNA synthesis; however, in combination with Twinkle, single strands of DNA up to 2 kb can be synthesized. When ssDNA-binding protein (mtSSB) is added to mix, DNA products up to about 16 kb could be made—i.e., the same size as mammalian mtDNA.
For the sake of argument, let's suppose a scientist sets upon the task of trying to build all the chromosomes from just sequence and epigenetic tag information. Is this even possible in theory? In other words, is there enough information in the raw code to insure that the nucleosomes are provisioned in the right spots and loops are extruded to the right lengths, and so on; or would some functioning template of existing chromosomes to configure be required to properly constrain construction of another one?
Stated another way, is there just one stable or optimal solution for configuring a functioning chromosome from scratch (like successful proteins always folding in the proper way), or are there so many possible solutions that completely different or even nonfunctional organisms would result? It would seem that there would be many ways. These are not simply idle speculations, because many would-be creature creators are now at their genetic drawing boards. The difference between an elephant and a mammoth appears to be largely one of sequence and epigenetics. Therefore, reconstructing mammoths from degraded genetic sources is probably not such a stretch.
But what about making a dragon? Considering that we will soon have fairly complete chromosome sequence and structural information for all extant animals, and even extinct pterosaurs and tyrannosaurids, it might not be entirely inconceivable that a good BioCAD package could be used to develop a reasonable facsimile of a dragon. And yet, there are no dragons in the fossil record. At least on Earth, a dragon is a Garden of Eden animal—there is no way to get one from scratch, presumably. It seems reasonable that a dragon could be a viable physiologic form in the sense that if a large pterosaur was surgically modified into a suitable dragon, it could persist, albeit likely sterile, for some time. It is another question entirely whether such an animal could ever be coded in DNA and clothed into stable chromosomes that could develop into a living, reproducing animal.
While the problem of building a set of viable dragon chromsomes is a step beyond the genome-in-a-box effort for existing animal or even human chromsomes, it highlights a potential shortcoming of the scheme. It may be possible to eventually make a certain chromsome from sequence, but perhaps not possible to make the actually correct chromosome, particularly if it is done in isolation. In other words, the chromosomes may need each other, physically, to build themselves. Chromosomes interact differently with each other at different phases of the cell cycle, and in various stages of development. Distinct phases of epigenetic reboot take place during germ cell and embryonic development.
For example, differentiating sperm cells cells shed their histones and temporarily don protamine coats to adopt a more compact configuration in preparation for meeting their counterparts in the egg. For some time thereafter, chromosomes retain some memory of their original parent. Since chromosomes continually change their own structure over the course of cell and organism development, a ground-level question for a 'genome in a box' might be: Which stage chromosome do we want to try to make first?
More information: Anthony Birnie et al. Genome-in-a-Box: Building a Chromosome from the Bottom Up, ACS Nano (2020). DOI: 10.1021/acsnano.0c07397
Journal information: ACS Nano